PRKAG1 Primary Antibody
Item Information
Catalog #
Size
Price
Description
The protein encoded by this gene is a regulatory subunit of the AMP-activated protein kinase (AMPK). AMPK is a heterotrimer consisting of an alpha catalytic subunit, and non-catalytic beta and gamma subunits. AMPK is an important energy-sensing enzyme that monitors cellular energy status. In response to cellular metabolic stresses, AMPK is activated, and thus phosphorylates and inactivates acetyl-CoA carboxylase (ACC) and beta-hydroxy beta-methylglutaryl-CoA reductase (HMGCR), key enzymes involved in regulating de novo biosynthesis of fatty acid and cholesterol. This subunit is one of the gamma regulatory subunits of AMPK. Alternatively spliced transcript variants encoding distinct isoforms have been observed.
Product Overview
Entrez GenelD
5571
Aliases
AMPKG
Clone#
7H3E9
Host / Isotype
Mouse / IgG1
Species Reactivity
Human
Immunogen
Purified recombinant fragment of human PRKAG1 (AA: 230-331) expressed in E. Coli.
Formulation
Purified antibody in PBS with 0.05% sodium azide.
Storage
Store at 4°C short term. Aliquot and store at -20°C long term. Avoid freeze/thaw cycles.
Product Applications
WB (Western Blot)
1/500 - 1/2000
IHC_P(Immunohistochemistry)
1/200 - 1/1000
ICC (Immunocytochemistry)
1/200 - 1/1000
FCM (Flow Cytometry)
1/200 - 1/400
ELISA
1/10000
References
1. Circ Res. 2012 Aug 31;111(6):800-14.
2. Circ Res. 2012 Apr 27;110(9):1192-201.
2. Circ Res. 2012 Apr 27;110(9):1192-201.
Product Image
Western Blot

Figure 1: Western blot analysis using PRKAG1 mAb against human PRKAG1 (AA: 230-331) recombinant protein. (Expected MW is 37.4 kDa)
Western Blot

Figure 2: Western blot analysis using PRKAG1 mAb against HEK293 (1) and PRKAG1 (AA: 230-331)-hIgGFc transfected HEK293 (2) cell lysate.
Immunofluorescence analysis

Figure 3: Immunofluorescence analysis of HepG2 cells using PRKAG1 mouse mAb (green). Blue: DRAQ5 fluorescent DNA dye. Secondary antibody from Fisher (Cat#: 35503)
Flow cytometric

Figure 4: Flow cytometric analysis of HepG2 cells using PRKAG1 mouse mAb (green) and negative control (red).
Immunohistochemical analysis

Figure 5: Immunohistochemical analysis of paraffin-embedded prostate cancer tissues using PRKAG1 mouse mAb with DAB staining.
Immunohistochemical analysis

Figure 6: Immunohistochemical analysis of paraffin-embedded kidney tissues using PRKAG1 mouse mAb with DAB staining.
Elisa
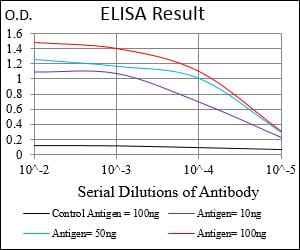
Black line: Control Antigen (100 ng); Purple line: Antigen(10ng); Blue line: Antigen (50 ng); Red line: Antigen (100 ng);
For Research Use Only. Not for use in diagnostic procedures.

